AI Cracks Plant DNA Code: Language Models Poised to Revolutionize Genomics and Agriculture
AI Cracks Plant DNA Code: Language Models Poised to Revolutionize Genomics and Agriculture
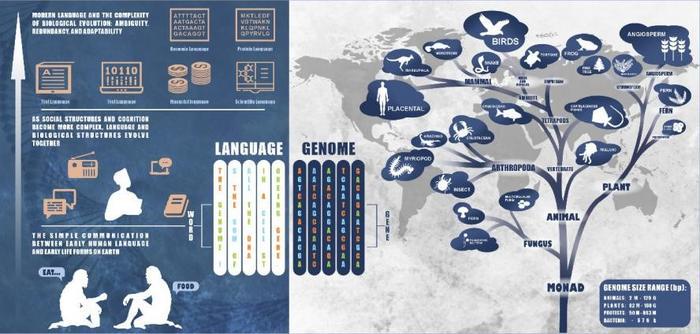
In a groundbreaking advancement at the nexus of artificial intelligence and plant biology, a new study spearheaded by Meiling Zou, Haiwei Chai, and Zhiqiang Xia from Hainan University heralds a transformative era in plant genomics research. By harnessing the power of large language models (LLMs)—AI architectures originally designed for human language processing—scientists are now unveiling the intricate lexicon embedded in plant genomes. This pioneering work, published in the journal Tropical Plants, details how these AI-driven models decode the complex language of genetic sequences to unlock unprecedented biological insights and propel agricultural innovation.
Historically, the domain of plant genomics has stumbled over the colossal complexity intrinsic to plant DNA. Vast, variable, and often poorly annotated datasets pose significant challenges for traditional machine learning techniques, which require large volumes of high-quality labeled data. Unlike human languages, which are rich in structured grammar and semantics, genomic sequences represent a fundamentally different modality of biological information—strings of nucleotides whose regulatory and functional elements reflect sophisticated hierarchical patterns. The recent study confronts this challenge by reimagining genome sequences as a language-like system, thus enabling large language models to process and predict genetic functions with remarkable accuracy.
The crux of this research lies in recognizing the striking structural parallels between natural language and genomic codes. DNA can be conceptualized as a sequence of “words” composed of nucleotide letters—adenine, thymine, cytosine, and guanine—that combine to form meaningful “sentences” or motifs regulating gene expression and cellular function. By training LLMs on massive datasets of plant genomic sequences, the researchers have demonstrated that these models can learn to identify complex features such as promoters, enhancers, and other regulatory elements that orchestrate gene activity across various tissues and developmental stages.
The study explores the performance of multiple LLM architectures specifically tailored for plant genomic analysis. Encoder-only models, exemplified by DNABERT, focus on interpreting input sequences to extract meaningful representations. Decoder-only models like DNAGPT facilitate generative tasks, predicting downstream sequence patterns or functional annotations. Additionally, encoder-decoder hybrids such as ENBED enable bidirectional understanding and prediction, enhancing model versatility. The researchers employed a rigorous methodology involving initial pre-training on expansive raw genomic data, followed by fine-tuning
AI Unraveling Plant DNA: Language Models Poised to Revolutionize Genomics and Agriculture
Next PostMagnetic Soft Robot Innovates Intelligent Bladder Control
Wheat Extreme Dwarfism Disrupts Gluten Composition and Compromises Baking Quality