Revolutionizing High-Speed Dynamic Fluorescence Imaging with Deep Learning-Enhanced Denoising Techniques
Revolutionizing High-Speed Dynamic Fluorescence Imaging with Deep Learning-Enhanced Denoising Techniques
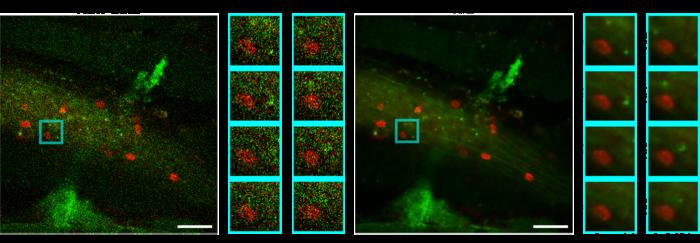
Researchers have unveiled a groundbreaking advancement in the field of fluorescence microscopy, designed to resolve one of the most significant challenges facing scientists: image degradation due to noise in dynamic in vivo imaging. This innovative method, recently published in the journal PhotoniX, presents a self-supervised deep learning approach known as Temporal-gradient empowered Denoising (TeD). The newly developed technique stands to revolutionize the way researchers capture and analyze high-speed biological processes, which are often severely obscured by noise.
Fluorescence microscopy is a powerful tool used to explore live biological processes at both the cellular and subcellular levels. However, its application is frequently limited by low signal-to-noise ratios, especially under photon-limited conditions where crucial biological signals can become faint and indistinguishable. This situation necessitates rapid imaging techniques that can lead to significant image degradation due to various types of noise. The introduction of the TeD model promises to change that dynamic completely, as it claims to enhance the quality of fluorescence images without needing pristine reference images for training purposes.
A notable feature of the TeD model is its incorporation of a temporal gradient-based attention mechanism. This mechanism adeptly detects spatial motion and adjusts the use of temporal redundancy for denoising processes. Essentially, the model makes intelligent decisions about how to filter out noise while preserving the integrity of the biological signals being observed. Researchers are able to process time-lapse sequences using this approach, allowing for the selective utilization of only the most relevant spatiotemporal features. This careful and deliberate handling of data allows for the preservation of moving structures like circulating blood cells, among others.
Validation of the TeD model has been conducted across multiple imaging modalities, including both confocal and two-photon fluorescence microscopy. The results have been promising, showcasing the model’s capacity to recover fine structural details that were previously difficult, if not impossible, to discern under static or dynamic conditions. A series of quantitative assessments have confirmed that TeD significantly enhances the signal-to-noise ratio and structural fidelity compared to traditional methods, setting a new standard in the field of imaging analysis.
The implications of this research extend far beyond just technical advancements. By enabling the collection of better quality fluorescence images under real-world in vivo conditions, researchers can delve deeper into the spatiotemporal dynamics of various biological processes. Such advancements could lead to new and unexpected findings in pathological research, granting scientists the tools they need to explore complex biological systems more effectively.
Furthermore, one of the most significant advantages of the TeD model is its applicability to real-world scenarios. Unlike conventional supervised models that rely on clean ground truth images for training, TeD is capable of operating effectively in environments where such ideal reference images are not available. This flexibility opens up a myriad of opportunities for progression in biological imaging, particularly in scenarios involving rapid biological dynamics.
The research team behind TeD, led by W. Lee et al., has underscored the importance of this breakthrough in advancing scientific understanding of dynamic biological processes. The ability to achieve high-resolution images of fluctuating biological phenomena can lead to new insights into cellular behaviors and interactions on a more intricate level. As a result, the TeD approach is poised to assist in a vast array of research areas, ranging from developmental biology to neuroscience and beyond.
Moreover, the publication of this study adds to the growing body of work emphasizing the importance of machine learning and deep learning techniques in scientific research. As technology continues to aid scientists in overcoming complex challenges, we can expect a ripple effect across numerous fields. These breakthroughs are not only significant for their immediate applications but also for the foundational advancements they build upon in the future of scientific endeavor.
The research community is already buzzing with anticipation for the potential applications of TeD in existing studies entangled with biological imaging. With significant improvements noted in both signal clarity and detail recovery, this method stands to enhance ongoing research efforts aimed at unraveling biological mysteries. As scientists begin to integrate TeD into their workflow, we can expect a shift in the design and execution of in vivo imaging experiments, fostering a new era of biological understanding.
The journey of TeD does not end here. As researchers continue to refine this model and explore its full potential, further studies are likely to emerge. The evaluations of TeD’s effectiveness across various disciplines will provide even deeper insights into how this model can be tailored to meet the demands of specific research questions. It may lead to the development of additional tools and methodologies that can capitalize on the successes of TeD, ensuring ongoing innovation.
Overall, the introduction of the TeD model marks a significant milestone in fluorescence microscopy and imaging analysis. The research not only provides a novel solution to a long-standing problem in the field but also opens new avenues for exploration and understanding of intricate biological systems. As scientists continue to apply and expand upon this innovative technology, we stand on the brink of uncovering new biological truths and advancing the frontiers of modern science.
Subject of Research: Animals
Article Title: Self-supervised denoising of dynamic fluorescence images via temporal gradient-empowered deep learning
News Publication Date: 23-May-2025
Web References: 10.1186/s43074-025-00173-8
References: None available
Image Credits: W. Lee et al. 2025 Springer Nature Publishing Group
Keywords
Denoising, fluorescence microscopy, temporal gradient, deep learning, imaging analysis, in vivo imaging, dynamic biological processes, signal-to-noise ratio, spatiotemporal dynamics.
Decades-Old Bladder Cancer Treatment Yields New Insights to Enhance Immunotherapy Advances
Next PostRevolutionary Sensor Technology Unveils 3D Structure of Sound Waves Triggered by Earthquakes
SFU Unveils Canada’s Fastest Academic Supercomputer Following $80 Million Upgrade